# 1. Mission Setup
To begin with, we have to load libraries. Be certain all of them are correctly put in. Additionally, I’m utilizing Python 3.12.3.
## 1.1 Load libraries# If it is advisable to set up any library, run beneath:
# pip set up library1 library2 library3 ...
# Primary Libraries
import os # For file operations
import gc # For rubbish assortment, it avoids RAM reminiscence points
import numpy as np # For coping with matrices
import pandas as pd # For coping with dataframes
import janitor # For information cleansing (primarily column names)
import numexpr # For quick pd.question() manipulation
import inflection # For string manipulation
import unidecode # For string manipulation
# Geospatial Libraries
import geopandas as gpd # For coping with shapefiles
import pyogrio # For quick .gpkg file manipulation
import ee # For Google Earth Engine API
import contextily as ctx # For basemaps
import folium # For interactive maps
# Shapely Objects and Geometry Manipulation
from shapely.geometry import mapping, Polygon, Level, MultiPolygon, LineString # For geometry manipulation
# Raster Information Manipulation and Visualization
import rasterio # For raster manipulation
from rasterio.masks import masks # For raster information manipulation
from rasterio.plot import present # For raster information visualization
# Plotting and Visualization
import matplotlib.pyplot as plt # For plotting and information visualization
from matplotlib.colours import ListedColormap, Normalize # For colour manipulation
import matplotlib.colours as colours # For colour manipulation
import matplotlib.patches as mpatches # For creating patch objects
import matplotlib.cm as cm # For colormaps
# Information Storage and Manipulation
import pyarrow # For environment friendly information storage and manipulation
# Video Making
from moviepy.editor import ImageSequenceClip # For creating movies (part 4.7 solely) - examine this in case you have points: https://github.com/kkroening/ffmpeg-python
Then, be sure to have a folder for this undertaking. All assets and outputs will probably be saved there. This folder could be situated in your native drive, a cloud-based storage resolution, or in a selected folder on Google Drive the place you’ll save the rasters retrieved utilizing the GEE API.
When working your code, be sure that to vary the deal with beneath to your undertaking path. Home windows customers, all the time keep in mind to make use of as a substitute of /.
# 1.2 Set working listing
project_path = 'path_to_your_project_folder' # The place you'll save all outcomes and assets should be in
os.chdir(project_path) # All assets on the undertaking are relative to this path# 1.3 Additional settings
pd.set_option('compute.use_numexpr', True) # Use numexpr for quick pd.question() manipulation
Lastly, this operate is helpful for plotting geometries over OpenStreetMap (OSM). It’s significantly useful when working with unknown shapefiles to make sure accuracy and keep away from errors.
## 1.4 Set operate to plot geometries over an OSM
def plot_geometries_on_osm(geometries, zoom_start=10):# Calculate the centroid of all geometries to middle the map
centroids = [geometry.centroid for geometry in geometries]
avg_x = sum(centroid.x for centroid in centroids) / len(centroids)
avg_y = sum(centroid.y for centroid in centroids) / len(centroids)
# Create a folium map centered across the common centroid
map = folium.Map(location=[avg_y, avg_x], zoom_start=zoom_start)
# Add every geometry to the map
for geometry in geometries:
geojson = mapping(geometry) # Convert the geometry to GeoJSON
folium.GeoJson(geojson).add_to(map)
return map
# 2. Single Instance: Acrelândia (AC) in 2022
For instance to create instinct of the method, we are going to save, clear, and plot land use in Acrelândia (AC) in 2022. It’s a metropolis in the course of the AMACRO area (the three-state border of Amazonas, Acre, and Rondônia), the place the usually untouched forest is being quickly destroyed.
On this part, I’ll clarify step-by-step of the script, after which standardize the method to run it for a number of locations over a number of years. Since saving giant rasters utilizing the API could be a gradual course of, I like to recommend utilizing it provided that it is advisable to cope with a couple of or small areas for a couple of years. Giant areas might take hours to save lots of on Google Drive, so I like to recommend downloading the heavy LULC recordsdata for the entire nation after which cleansing them, as we are going to do in a future publish.
To run the code, first obtain and save IBGE’s¹ Brazilian cities shapefiles (choose Brazil > Municipalities). Bear in mind, you should utilize any shapefile in Brazil to carry out this algorithm.
## 2.1 Get the geometry of the world of curiosity (Acrelândia, AC)
brazilian_municipalities = gpd.read_file('municipios/file.shp', engine='pyogrio', use_arrow=True) # Learn the shapefile - you should utilize some other shapefile right here. Shapefiles should be in your undertaking folder, as set in 1.2
brazilian_municipalities = brazilian_municipalities.clean_names() # Clear the column names (take away particular characters, areas, and so forth.)
brazilian_municipalities.crs = 'EPSG:4326' # Set the CRS to WGS84 (MapBiomas makes use of this CRS)
brazilian_municipalities
## 2.2 Get geometry for Acrelândia, AC
metropolis = brazilian_municipalities.question('nm_mun == "Acrelândia"') # Filter the geometry for Acrelândia, AC (could be some other metropolis or set of cities)
city_geom = metropolis.geometry.iloc[0] # Get the geometry of Acrelândia, AC
city_geom # See the geometry form
As soon as we’ve the shapefile we need to research correctly saved, we are going to create a bounding field round it to crop the MapBiomas full raster. Then, we are going to put it aside the GEE Python API.
## 2.3 Set the bounding field (bbox) for the world of curiosity
bbox = city_geom.bounds # Get the bounding field of the geometry
bbox = Polygon([(bbox[0], bbox[1]), (bbox[0], bbox[3]), (bbox[2], bbox[3]), (bbox[2], bbox[1])]) # Convert the bounding field to a Polygonbbox_xmin = bbox.bounds[0] # Get the minimal x coordinate of the bounding field
bbox_ymin = bbox.bounds[1] # Get the minimal y coordinate of the bounding field
bbox_xmax = bbox.bounds[2] # Get the utmost x coordinate of the bounding field
bbox_ymax = bbox.bounds[3] # Get the utmost y coordinate of the bounding field
bbox # See bbox round Acrelândia form
# Plot the bounding field and the geometry of Acrelandia over an OSM map
plot_geometries_on_osm([bbox, city_geom], zoom_start=10)
Now, we are going to entry the MapBiomas Google Earth Engine API. First, we have to create a cloud undertaking on GEE utilizing a Google Account. Be sure you have sufficient area in your Google Drive account.
Then, we have to authenticate the GEE Python API (solely as soon as). If you’re a VSCode person, discover that the token insertion field seems within the prime proper nook of the IDE.
All photographs from the MapBiomas LULC Assortment can be found in the identical asset. Discover that you could barely modify this script to work with different belongings within the GEE catalog and different MapBiomas collections.
## 2.4 Acess MapBiomas Assortment 8.0 utilizing GEE API
# import ee - already imported at 1.1ee.Authenticate() # Just for the primary time
ee.Initialize() # Run it each time you begin a brand new session
# Outline the MapBiomas Assortment 8.0 asset ID - retrieved from https://brasil.mapbiomas.org/en/colecoes-mapbiomas/
mapbiomas_asset = 'tasks/mapbiomas-workspace/public/collection8/mapbiomas_collection80_integration_v1'
asset_properties = ee.information.getAsset(mapbiomas_asset) # Examine the asset's properties
asset_properties # See properties
Right here, every band represents the LULC information for a given 12 months. Guarantee that the code beneath is correctly written. This selects the picture for the specified 12 months after which crops the uncooked raster for a bounding field across the area of curiosity (ROI) — Acrelândia, AC.
## 2.5 Filter the gathering for 2022 and crop the gathering to a bbox round Acrelândia, AC
12 months = 2022
band_id = f'classification_{12 months}' # bands (or yearly rasters) are named as classification_1985, classification_1986, ..., classification_2022mapbiomas_image = ee.Picture(mapbiomas_asset) # Get the pictures of MapBiomas Assortment 8.0
mapbiomas2022 = mapbiomas_image.choose(band_id) # Choose the picture for 2022
roi = ee.Geometry.Rectangle([bbox_xmin, bbox_ymin, bbox_xmax, bbox_ymax]) # Set the Area of Curiosity (ROI) to the bbox round Acrelândia, AC - set in 2.3
image_roi = mapbiomas2022.clip(roi) # Crop the picture to the ROI
Now, we save the cropped raster on Google Drive (in my case, into the ‘tutorial_mapbiomas_gee’ folder). Be sure you have created the vacation spot folder in your drive earlier than working.
I attempted to put it aside regionally, but it surely seems to be like it is advisable to save GEE rasters at Google Drive (let me know if you know the way to do it regionally). That is probably the most time-consuming a part of the code. For giant ROIs, this would possibly take hours. Examine your GEE activity supervisor to see if the rasters have been correctly loaded to the vacation spot folder.
## 2.6 Export it to your Google Drive (guarantee you might have area there and that it's correctly arrange)# Obs 1: Recall it is advisable to authenticate the session, because it was carried out on 2.4
# Obs 2: Guarantee you might have sufficient area on Google Drive. Free model solely provides 15 Gb of storage.
export_task = ee.batch.Export.picture.toDrive(
picture=image_roi, # Picture to export to Google Drive as a GeoTIFF
description='clipped_mapbiomas_collection8_acrelandia_ac_2022', # Activity description
folder='tutorial_mapbiomas_gee', # Change this to the folder in your Google Drive the place you need to save the file
fileNamePrefix='acrelandia_ac_2022', # File identify (change it if you wish to)
area=roi.getInfo()['coordinates'], # Area to export the picture
scale=30,
fileFormat='GeoTIFF'
)
# Begin the export activity
export_task.begin()
# 3. Plot the Map
Now we’ve a raster with LULC information for a bounding field round Acrelândia in 2022. That is saved on the deal with beneath (at Google Drive). First, let’s see the way it seems to be.
## 3.1 Plot the orginal raster over a OSM
file_path = 'path_of_exported_file_at_google_drive.tif' # Change this to the trail of the exported file# Plot information
with rasterio.open(file_path) as src:
information = src.learn(1)
print(src.meta)
print(src.crs)
present(information)
In MapBiomas LULC Assortment 8, every pixel represents a selected land use sort in accordance with this listing. As an illustration, ‘3’ means ‘Pure Forest’, ‘15’ means ‘Pasture’, and ‘0’ means ‘No information’ (pixels within the raster not inside the Brazilian borders).
We are going to discover the info we’ve earlier than plotting it.
## 3.2 See distinctive values
unique_values = np.distinctive(information)
unique_values # Returns distinctive pixels values within the raster# 0 = no information, components of the picture exterior Brazil
## 3.3 See the frequency of every class (besides 0 - no information)
unique_values, counts = np.distinctive(information[data != 0], return_counts=True) # Get the distinctive values and their counts (besides zero)
pixel_counts = pd.DataFrame({'worth': unique_values, 'rely': counts})
pixel_counts['share'] = (pixel_counts['count'] / pixel_counts['count'].sum())*100
pixel_counts
On the finish of the day, we’re working with a big matrix wherein every ingredient represents how every tiny 30m x 30m piece of land is used.
## 3.4 See the precise raster (a matrix wherein every ingredient represents a pixel worth - land use code on this case)
information
Now, we have to manage our raster information. As a substitute of categorizing every pixel by precise land use, we are going to categorize them extra broadly. We are going to divide pixels into pure forest, pure non-forest vegetation, water, pasture, agriculture, and different makes use of. Particularly, we’re enthusiastic about monitoring the conversion of pure forest to pasture. To realize this, we are going to reassign pixel values based mostly on the mapbiomas_categories dict beneath, which follows with MapBiomas’ land use and land cowl (LULC) categorization.
The code beneath crops the raster to Acrelândia’s limits and reassigns pixels in accordance with the mapbiomas_categories dict. Then, it saves it as a brand new raster at ‘reassigned_raster_path’. Observe that the previous raster was saved on Google Drive (after being downloaded utilizing GEE’s API), whereas the brand new one will probably be saved within the undertaking folder (in my case, a OneDrive folder on my PC, as set in part 1.2). From right here onwards, we are going to use solely the reassigned raster to plot the info.
That is the principle a part of the script. In case you have doubts about what is going on right here (cropping for Acrelândia after which reassigning pixels to broader classes), I like to recommend working it and printing outcomes for each step.
mapbiomas_categories = {
# Forest (= 3)
1:3, 3:3, 4:3, 5:3, 6:3, 49:3, # That's, values 1, 3, 4, 5, 6, and 49 will probably be reassigned to three (Forest)
# Different Non-Forest Pure Vegetation (= 10)
10:10, 11:10, 12:10, 32:10, 29:10, 50:10, 13:10, # That's, values 10, 11, 12, 32, 29, 50, and 13 will probably be reassigned to 10 (Different Non-Forest Pure Vegetation)
# Pasture (= 15)
15:15,
# Agriculture (= 18)
18:18, 19:18, 39:18, 20:18, 40:18, 62:18, 41:18, 36:18, 46:18, 47:18, 35:18, 48:18, 21:18, 14:18, 9:18, # That's, values 18, 19, 39, 20, 40, 62, 41, 36, 46, 47, 35, 48, 21, 14, and 9 will probably be reassigned to 18 (Agriculture)
# Water ( = 26)
26:26, 33:26, 31:26, # That's, values 26, 33, and 31 will probably be reassigned to 26 (Water)
# Different (= 22)
22:22, 23:22, 24:22, 30:22, 25:22, 27:22, # That's, values 22, 23, 24, 30, 25, and 27 will probably be reassigned to 22 (Different)
# No information (= 255)
0:255 # That's, values 0 will probably be reassigned to 255 (No information)
}
## 3.5 Reassing pixels values to the MapBiomas customized common classes and crop it to Acrelandia, AC limits
original_raster_path = 'path_to_your_google_drive/tutorial_mapbiomas_gee/acrelandia_ac_2022.tif'
reassigned_raster_path = 'path_to_reassigned_raster_at_project_folder' # Someplace within the undertaking folder set at 1.2with rasterio.open(original_raster_path) as src:
raster_array = src.learn(1)
out_meta = src.meta.copy() # Get metadata from the unique raster
# 3.5.1. Crop (or masks) the raster to the geometry of city_geom (on this case, Acrelandia, AC) and thus take away pixels exterior the town limits (will probably be assigned to no information = 255)
out_image, out_transform = rasterio.masks.masks(src, [city_geom], crop=True)
out_meta.replace({
"top": out_image.form[1],
"width": out_image.form[2],
"rework": out_transform
}) # Replace metadata to the brand new raster
raster_array = out_image[0] # Get the masked raster
modified_raster = np.zeros_like(raster_array) # Base raster stuffed with zeros to be modified
# 3.5.2. Reassign every pixel based mostly on the mapbiomas_categories dictionary
for original_value, new_value in mapbiomas_categories.gadgets():
masks = (raster_array == original_value) # Create a boolean masks for the unique worth (True = Substitute, False = Do not change)
modified_raster[mask] = new_value # Substitute the unique values with the brand new values, when wanted (that's, when the masks is True)
out_meta = src.meta.copy() # Get metadata from the unique raster
out_meta.replace(dtype=rasterio.uint8, rely=1) # Replace metadata to the brand new raster
with rasterio.open(reassigned_raster_path, 'w', **out_meta) as dst: # Write the modified raster to a brand new file on the reassigned_raster_path
dst.write(modified_raster.astype(rasterio.uint8), 1)
## 3.6 See the frequency of pixels within the reassigned raster
with rasterio.open(reassigned_raster_path) as src:
raster_data = src.learn(1)
unique_values = np.distinctive(raster_data)
total_non_zero = np.sum(raster_data != 255) # Rely the entire variety of non-zero pixelsfor worth in unique_values:
if worth != 255: # Exclude no information (= 255)
rely = np.sum(raster_data == worth) # Rely the variety of pixels with the worth
share = rely / total_non_zero # Calculate the share of the worth
share = share.spherical(3) # Spherical to three decimal locations
print(f"Worth: {worth}, Rely: {rely}, Share: {share}")
Now we plot the info with generic colours. We are going to improve the map later, however that is only a first (or second?) look. Discover that I particularly set 255 (= no information, pixels exterior Acrelândia) to be white for higher visualization.
## 3.7 Plot the reassigned raster with generic colours
with rasterio.open(reassigned_raster_path) as src:
information = src.learn(1) # Learn the raster information
unique_values = np.distinctive(information) # Get the distinctive values within the rasterplt.determine(figsize=(10, 8)) # Set the determine dimension
cmap = plt.cm.viridis # Utilizing Viridis colormap
norm = Normalize(vmin=information.min(), vmax=26) # Normalize the info to the vary of the colormap (max = 26, water)
masked_data = np.ma.masked_where(information == 255, information) # Masks no information values (255)
plt.imshow(masked_data, cmap=cmap, norm=norm) # Plot the info with the colormap
plt.colorbar(label='Worth') # Add a colorbar with the values
plt.present()
Now it’s time to create an attractive map. I’ve chosen Matplotlib as a result of I would like static maps. In case you desire interactive choropleths, you should utilize Plotly.
For additional particulars on choropleths with Matplotlib, examine its documentation, GeoPandas information, and the good Yan Holtz’s Python Graph Gallery — the place I get a lot of the inspiration and instruments for DataViz in Python. Additionally, for lovely colour palettes, coolors.co is a superb useful resource.
Be sure you have all information visualization libraries correctly loaded to run the code beneath. I additionally tried to vary the order of patches, however I didn’t know how you can. Let me know if you happen to learn the way to do it.
## 3.8 Plot the reassigned raster with customized colours# Outline the colours for every class - discover it is advisable to comply with the identical order because the values and should be numerically growing or reducing (nonetheless must learn the way to resolve it)
values = [3, 10, 15, 18, 22, 26, 255] # Values to be coloured
colors_list = ['#6a994e', '#a7c957', '#c32f27', '#dda15e', '#6c757d', '#0077b6','#FFFFFF'] # HEX codes of the colours used
labels = ['Natural Forest', 'Other Natural Vegetation', 'Pasture', 'Agriculture', 'Others', 'Water', 'No data'] # Labels displayed on the legend
cmap = colours.ListedColormap(colors_list) # Create a colormap (cmap) with the colours
bounds = values + [256] # Add a price to the tip of the listing to incorporate the final colour
norm = colours.BoundaryNorm(bounds, cmap.N) # Normalize the colormap to the values
img = plt.imshow(raster_data, interpolation='nearest', cmap=cmap, norm=norm) # Plot the info with the colormap
legend_patches = [mpatches.Patch(color=colors_list[i], label=labels[i]) for i in vary(len(values)-1)] # Create the legend patches withou the final one (255 = no information)
# Create the legend
plt.legend(handles = legend_patches, # Add the legend patches
bbox_to_anchor = (0.5, -0.02), # Place the legend beneath the plot
loc = 'higher middle', # Place the legend within the higher middle
ncol = 3, # Variety of columns
fontsize = 9, # Font dimension
handlelength=1,# Size of the legend handles
frameon=False) # Take away the body across the legend
plt.axis('off') # Take away the axis
plt.title('Land Use in Acrelândia, AC (2022)', fontsize=20) # Add title
plt.savefig('figures/acrelandia_ac_2022.pdf', format='pdf', dpi=1800) # Put it aside as a PDF on the figures folder
plt.present()
4. Standardized Features
Now that we’ve constructed instinct on how you can obtain, save, clear, and plot MapBiomas LULC rasters. It’s time to generalize the method.
On this part, we are going to outline capabilities to automate these steps for any form and any 12 months. Then, we are going to execute these capabilities in a loop to investigate a selected metropolis — Porto Acre, AC — from 1985 to 2022. Lastly, we are going to make a video illustrating the LULC evolution in that space over the required interval.
First, save a bounding field (bbox) across the Area of Curiosity (ROI). You solely must enter the specified geometry and specify the 12 months. This operate will save the bbox raster across the ROI to your Google Drive.
## 4.1 For a generic geometry in any 12 months, save a bbox across the geometry to Google Drivedef get_mapbiomas_lulc_raster(geom, geom_name, 12 months, folder_in_google_drive):
ee.Authenticate() # Just for the primary time
ee.Initialize() # Run it each time you begin a brand new session
my_geom = geom
bbox = my_geom.bounds # Get the bounding field of the geometry
bbox = Polygon([(bbox[0], bbox[1]), (bbox[0], bbox[3]), (bbox[2], bbox[3]), (bbox[2], bbox[1])]) # Convert the bounding field to a Polygon
bbox_xmin = bbox.bounds[0] # Get the minimal x coordinate of the bounding field
bbox_ymin = bbox.bounds[1] # Get the minimal y coordinate of the bounding field
bbox_xmax = bbox.bounds[2] # Get the utmost x coordinate of the bounding field
bbox_ymax = bbox.bounds[3] # Get the utmost y coordinate of the bounding field
mapbiomas_asset = 'tasks/mapbiomas-workspace/public/collection8/mapbiomas_collection80_integration_v1'
band_id = f'classification_{12 months}'
mapbiomas_image = ee.Picture(mapbiomas_asset) # Get the pictures of MapBiomas Assortment 8.0
mapbiomas_data = mapbiomas_image.choose(band_id) # Choose the picture for 2022
roi = ee.Geometry.Rectangle([bbox_xmin, bbox_ymin, bbox_xmax, bbox_ymax]) # Set the Area of Curiosity (ROI) to the bbox across the desired geometry
image_roi = mapbiomas_data.clip(roi) # Crop the picture to the ROI
export_task = ee.batch.Export.picture.toDrive(
picture=image_roi, # Picture to export to Google Drive as a GeoTIFF
description=f"save_bbox_around_{geom_name}_in_{12 months}", # Activity description
folder=folder_in_google_drive, # Change this to the folder in your Google Drive the place you need to save the file
fileNamePrefix=f"{geom_name}_{12 months}", # File identify
area=roi.getInfo()['coordinates'], # Area to export the picture
scale=30,
fileFormat='GeoTIFF'
)
export_task.begin() # Discover that importing these rasters to Google Drive might take some time, specifically for giant areas
# Take a look at it utilizing Rio de Janeiro in 2022
folder_in_google_drive = 'tutorial_mapbiomas_gee'
rio_de_janeiro = brazilian_municipalities.question('nm_mun == "Rio de Janeiro"')
rio_de_janeiro.crs = 'EPSG:4326' # Set the CRS to WGS84 (this undertaking default one, change if wanted)
rio_de_janeiro_geom = rio_de_janeiro.geometry.iloc[0] # Get the geometry of Rio de Janeiro, RJteste1 = get_mapbiomas_lulc_raster(rio_de_janeiro_geom, 'rio_de_janeiro', 2022, folder_in_google_drive)
Second, crop the raster to incorporate solely the pixels inside the geometry and put it aside as a brand new raster.
I selected to put it aside on Google Drive, however you may change `reassigned_raster_path` to put it aside anyplace else. In case you change it, be sure that to replace the remainder of the code accordingly.
Additionally, you may reassign pixels as wanted by modifying the mapbiomas_categories dict. The left digit represents the unique pixel values, and the precise one represents the reassigned (new) values.
## 4.2 Crop the raster for the specified geometry
def crop_mapbiomas_lulc_raster(geom, geom_name, 12 months, folder_in_google_drive):
original_raster_path = f'path_to_your_google_drive/{folder_in_google_drive}/{geom_name}_{12 months}.tif'
reassigned_raster_path = f'path_to_your_google_drive/{folder_in_google_drive}/cropped_{geom_name}_{12 months}.tif'my_geom = geom
mapbiomas_categories = {
# Forest (= 3)
1:3, 3:3, 4:3, 5:3, 6:3, 49:3,
# Different Non-Forest Pure Vegetation (= 10)
10:10, 11:10, 12:10, 32:10, 29:10, 50:10, 13:10,
# Pasture (= 15)
15:15,
# Agriculture (= 18)
18:18, 19:18, 39:18, 20:18, 40:18, 62:18, 41:18, 36:18, 46:18, 47:18, 35:18, 48:18, 21:18, 14:18, 9:18,
# Water ( = 26)
26:26, 33:26, 31:26,
# Different (= 22)
22:22, 23:22, 24:22, 30:22, 25:22, 27:22,
# No information (= 255)
0:255
} # You'll be able to change this to no matter categorization you need, however simply keep in mind to adapt the colours and labels within the plot
with rasterio.open(original_raster_path) as src:
raster_array = src.learn(1)
out_meta = src.meta.copy() # Get metadata from the unique raster
# Crop the raster to the geometry of my_geom and thus take away pixels exterior the town limits (will probably be assigned to no information = 0)
out_image, out_transform = rasterio.masks.masks(src, [my_geom], crop=True)
out_meta.replace({
"top": out_image.form[1],
"width": out_image.form[2],
"rework": out_transform
}) # Replace metadata to the brand new raster
raster_array = out_image[0] # Get the masked raster
modified_raster = np.zeros_like(raster_array) # Base raster stuffed with zeros to be modified
# Reassign every pixel based mostly on the mapbiomas_categories dictionary
for original_value, new_value in mapbiomas_categories.gadgets():
masks = (raster_array == original_value) # Create a boolean masks for the unique worth (True = Substitute, False = Do not change)
modified_raster[mask] = new_value # Substitute the unique values with the brand new values, when wanted (that's, when the masks is True)
out_meta = src.meta.copy() # Get metadata from the unique raster
out_meta.replace(dtype=rasterio.uint8, rely=1) # Replace metadata to the brand new raster
with rasterio.open(reassigned_raster_path, 'w', **out_meta) as dst: # Write the modified raster to a brand new file on the reassigned_raster_path
dst.write(modified_raster.astype(rasterio.uint8), 1)
teste2 = crop_mapbiomas_lulc_raster(rio_de_janeiro_geom, 'rio_de_janeiro', 2022, folder_in_google_drive)
Now we see the frequency of every pixel within the cropped reassigned raster.
## 4.3 Plot the cropped raster
def pixel_freq_mapbiomas_lulc_raster(geom_name, 12 months, folder_in_google_drive):
reassigned_raster_path = f'path_to_your_google_drive/{folder_in_google_drive}/cropped_{geom_name}_{12 months}.tif'with rasterio.open(reassigned_raster_path) as src:
raster_data = src.learn(1)
unique_values = np.distinctive(raster_data)
total_non_zero = np.sum(raster_data != 255) # Rely the entire variety of non-zero pixels
for worth in unique_values:
if worth != 255: # Exclude no information (= 255)
rely = np.sum(raster_data == worth) # Rely the variety of pixels with the worth
share = rely / total_non_zero # Calculate the share of the worth
share = share.spherical(3)
print(f"Worth: {worth}, Rely: {rely}, Share: {share}")
teste3 = pixel_freq_mapbiomas_lulc_raster('rio_de_janeiro', 2022, folder_in_google_drive)
Lastly, we plot it on a map. You’ll be able to change the arguments beneath to regulate traits like colours, labels, legend place, font sizes, and so forth. Additionally, there’s an possibility to decide on the format wherein you need to save the info (often PDF or PNG). PDFs are heavier and protect decision, whereas PNGs are lighter however have decrease decision.
## 4.4 Plot the cropped raster
def plot_mapbiomas_lulc_raster(geom_name, 12 months, folder_in_google_drive,driver):
reassigned_raster_path = f'/Customers/vhpf/Library/CloudStorage/GoogleDrive-vh.pires03@gmail.com/My Drive/{folder_in_google_drive}/cropped_{geom_name}_{12 months}.tif'
with rasterio.open(reassigned_raster_path) as src:
raster_data = src.learn(1)# Outline the colours for every class - discover it is advisable to comply with the identical order because the values
values = [3, 10, 15, 18, 22, 26, 255] # Should be the identical of the mapbiomas_categories dictionary
colors_list = ['#6a994e', '#a7c957', '#c32f27', '#dda15e', '#6c757d', '#0077b6','#FFFFFF'] # Set your colours
labels = ['Natural Forest', 'Other Natural Vegetation', 'Pasture', 'Agriculture', 'Others', 'Water', 'No data'] # Set your labels
cmap = colours.ListedColormap(colors_list) # Create a colormap (cmap) with the colours
bounds = values + [256] # Add a price to the tip of the listing to incorporate the final colour
norm = colours.BoundaryNorm(bounds, cmap.N) # Normalize the colormap to the values
img = plt.imshow(raster_data, interpolation='nearest', cmap=cmap, norm=norm) # Plot the info with the colormap
legend_patches = [mpatches.Patch(color=colors_list[i], label=labels[i]) for i in vary(len(values)-1)] # Create the legend patches with out the final one (255 = no information)
# Create the legend
plt.legend(handles = legend_patches, # Add the legend patches
bbox_to_anchor = (0.5, -0.02), # Place the legend beneath the plot
loc = 'higher middle', # Place the legend within the higher middle
ncol = 3, # Variety of columns
fontsize = 9, # Font dimension
handlelength=1.5,# Size of the legend handles
frameon=False) # Take away the body across the legend
plt.axis('off') # Take away the axis
geom_name_title = inflection.titleize(geom_name)
plt.title(f'Land Use in {geom_name_title} ({12 months})', fontsize=20) # Add title
saving_path = f'figures/{geom_name}_{12 months}.{driver}'
plt.savefig(saving_path, format=driver, dpi=1800) # Put it aside as a .pdf or .png on the figures folder of your undertaking
plt.present()
teste4 = plot_mapbiomas_lulc_raster('rio_de_janeiro', 2022, folder_in_google_drive, 'png')
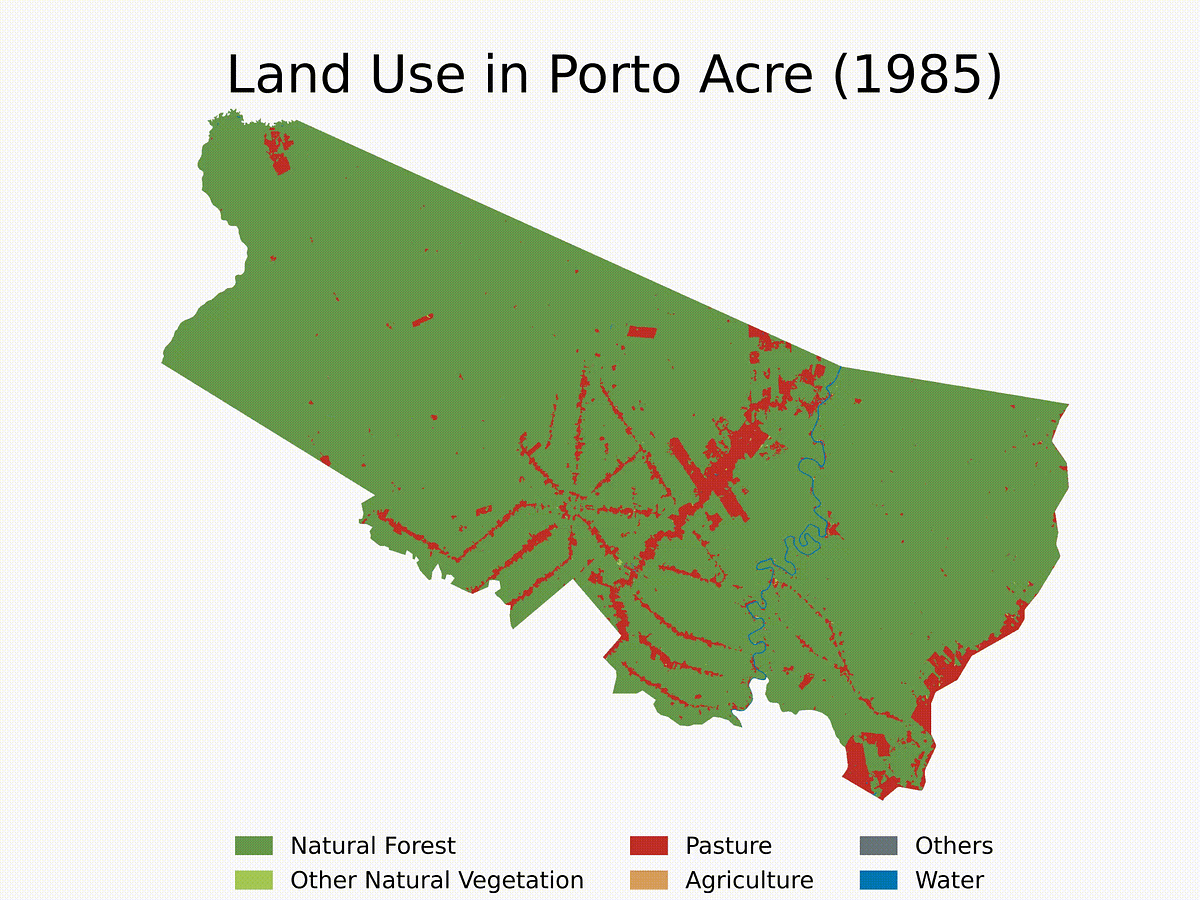
Lastly, right here’s an instance of how you can use the capabilities and create a loop to get the LULC evolution for Porto Acre (AC) since 1985. That’s one other metropolis within the AMACRO area with hovering deforestation charges.
## 4.5 Do it in only one operate - recall to save lots of rasters (4.1) earlier than
def clean_mapbiomas_lulc_raster(geom, geom_name, 12 months, folder_in_google_drive,driver):
crop_mapbiomas_lulc_raster(geom, geom_name, 12 months, folder_in_google_drive)
plot_mapbiomas_lulc_raster(geom_name, 12 months, folder_in_google_drive,driver)
print(f"MapBiomas LULC raster for {geom_name} in {12 months} cropped and plotted!")
## 4.6 Run it for a number of geometries for a number of years### 4.6.1 First, save rasters for a number of geometries and years
cities_list = ['Porto Acre'] # Cities to be analyzed - examine whether or not there are two cities in Brazil with the identical identify
years = vary(1985,2023) # Years to be analyzed (first 12 months in MapBiomas LULC == 1985)
brazilian_municipalities = gpd.read_file('municipios/file.shp', engine='pyogrio', use_arrow=True) # Learn the shapefile - you should utilize some other shapefile right here
brazilian_municipalities = brazilian_municipalities.clean_names()
brazilian_municipalities.crs = 'EPSG:4326' # Set the CRS to WGS84 (this undertaking default one, change if wanted)
selected_cities = brazilian_municipalities.question('nm_mun in @cities_list') # Filter the geometry for the chosen cities
selected_cities = selected_cities.reset_index(drop=True) # Reset the index
cities_ufs = [] # Create listing to append the complete names of the cities with their UF (state abbreviation, in portuguese)
nrows = len(selected_cities)
for i in vary(nrows):
metropolis = selected_cities.iloc[i]
city_name = metropolis['nm_mun']
city_uf = metropolis['sigla_uf']
cities_ufs.append(f"{city_name} - {city_uf}")
folder_in_google_drive = 'tutorial_mapbiomas_gee' # Folder in Google Drive to save lots of the rasters
for metropolis in cities_list:
for 12 months in years:
city_geom = selected_cities.question(f'nm_mun == "{metropolis}"').geometry.iloc[0] # Get the geometry of the town
geom_name = unidecode.unidecode(metropolis) # Take away latin-1 characters from the town identify - GEE doesn`t enable them
get_mapbiomas_lulc_raster(city_geom, geom_name, 12 months, folder_in_google_drive) # Run the operate for every metropolis and 12 months
### 4.6.2 Second, crop and plot the rasters for a number of geometries and years - Be sure you have sufficient area in your Google Drive and all rasters are there
for metropolis in cities_list:
for 12 months in years:
city_geom = selected_cities.question(f'nm_mun == "{metropolis}"').geometry.iloc[0]
geom_name = unidecode.unidecode(metropolis)
clean_mapbiomas_lulc_raster(city_geom, geom_name, 12 months, folder_in_google_drive,'png') # Run the operate for every metropolis and 12 months
gc.accumulate()
We are going to end the tutorial by creating a brief video displaying the evolution of deforestation within the municipality during the last 4 a long time. Observe that you could lengthen the evaluation to a number of cities and choose particular years for the evaluation. Be happy to customise the algorithm as wanted.
## 4.7 Make a clip with LULC evolution
img_folder = 'figures/porto_acre_lulc' # I created a folder to save lots of the pictures of the LULC evolution for Porto Acre inside project_path/figures
img_files = sorted([os.path.join(img_folder, f) for f in os.listdir(img_folder) if f.endswith('.png')]) # Will get all the pictures within the folder that finish with .png - be sure to solely have the specified photographs within the folderclip = ImageSequenceClip(img_files, fps=2) # 2 FPS, 0.5 second between frames
output_file = 'figures/clips/porto_acre_lulc.mp4' # Save clip on the clips folder
clip.write_videofile(output_file, codec='libx264') # It takes some time to create the video (3m30s in my laptop)



/cdn.vox-cdn.com/uploads/chorus_asset/file/25528444/1920x.jpg)



